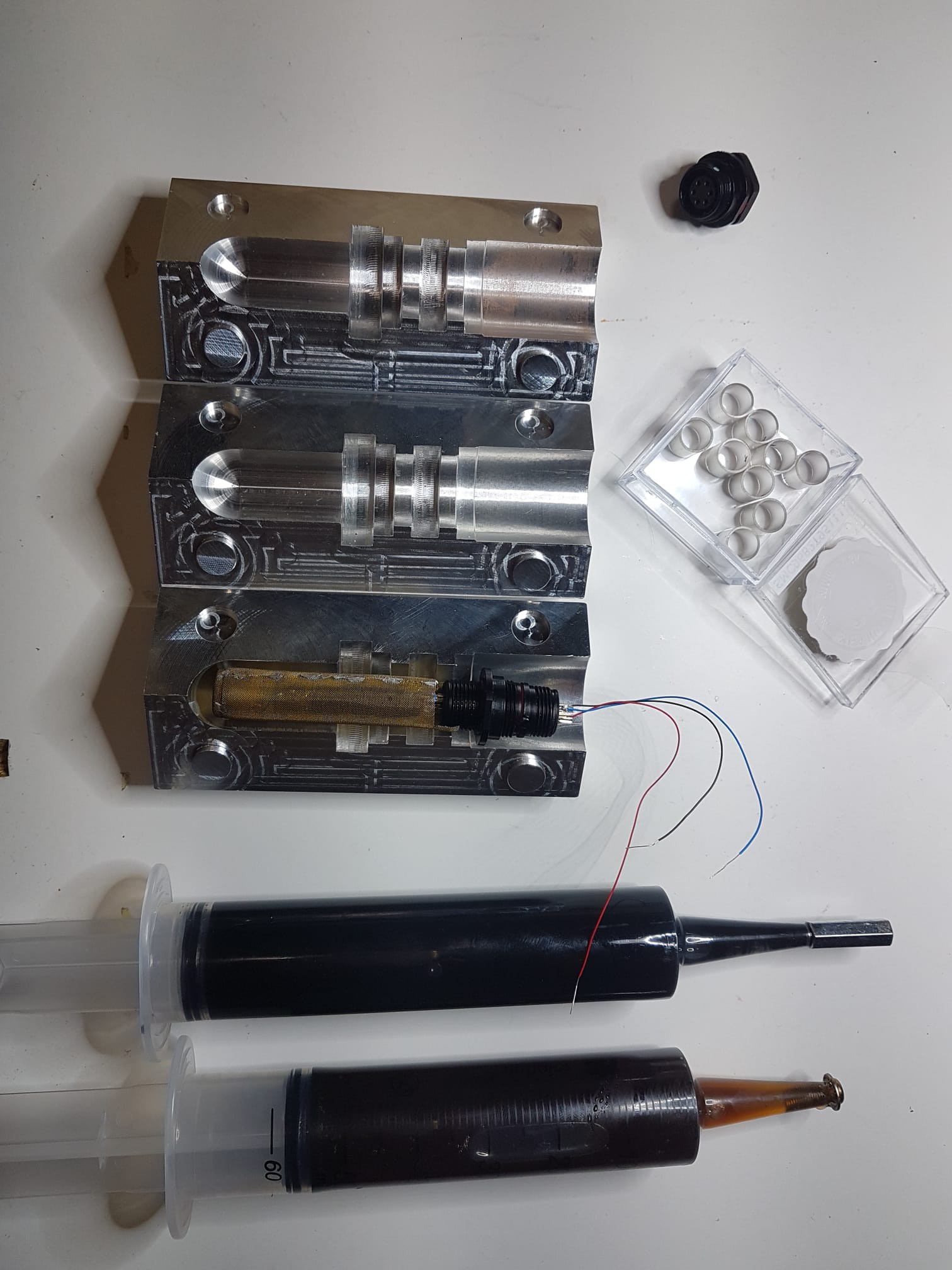
The core ingredients of this project are:
- Nvidia Jetson Nano development board / Raspberry Pi 4 + 4 GB RAM.
- Noctula fan nf a4x20 5v pwm (Nano only)
- ADC Pi shield for sensing battery and supply voltages.
- EVM3683-7-QN-01A Evaluation Board for supplying a steady 5v to the Nano.
- 5 Inch EDID Capacitive Touch Screen 800x480 HDMI Monitor TFT LCD Display.
- Dragino LoRa/GPS HAT for transmitting to the 'cloud' (Currently Pi 4 only)
- 12 V rechargeable battery pack.
- WavX bioacoustics R software package for wildlife acoustic feature extraction.
- Random Forest R classification software.
- In house developed deployment software.
- Full spectrum (384 kb per second) audio data.
- UltraMic 384 usb microphone.
- Waterproof case Max 004.
What have been the key challenges so far?
- Choosing the right software. Initially I started off using a package designed for music classification called ' PyAudioAnalysis' which gave options for both Random Forest and then human voice recognition Deep Learning using Tensorflow. Both systems worked ok, but the results were very poor. After some time chatting on this very friendly Facebook group: Bat Call Sound Analysis Workshop , I found a software package written in the R language with a decent tutorial that worked well within a few hours of tweaking. As a rule, if the tutorial is crap, then the software should probably be avoided! The same was true when creating the app with the touchscreen - I found one really good tutorial for GTK 3 + python, with examples, which set me up for a relatively smooth ride.
- After choosing to focus on detecting bats, finding quality bat data for my country. In theory, there should be numerous databases of full spectrum audio recordings in the UK and France, but when actually trying to download audio files, most of them seem to have been closed down or limited to the more obscure 'social calls'. The only option was to make my own recordings which entailed setting up the device overnight in my local nature reserves, by which I managed to find 7 species of bat. Undoubtedly, the data is the most important part of this project and I spent very many pleasant hours out in the wilderness with the detector and the sounds of these wonderful creatures.
- Using GTK 3 to produce the app. Whilst python itself is very well documented on Stack exchange etc, solving more detailed problems with GTK 3 was hard going. One bug was completely undocumented and took me 3 days to remove! The software is also rather clunky and not particularly user friendly or intuitive. Compared to ordinary programming with Python, GTK was NOT an enjoyable experience, although it's very rewarding to see the app in action.
![]()
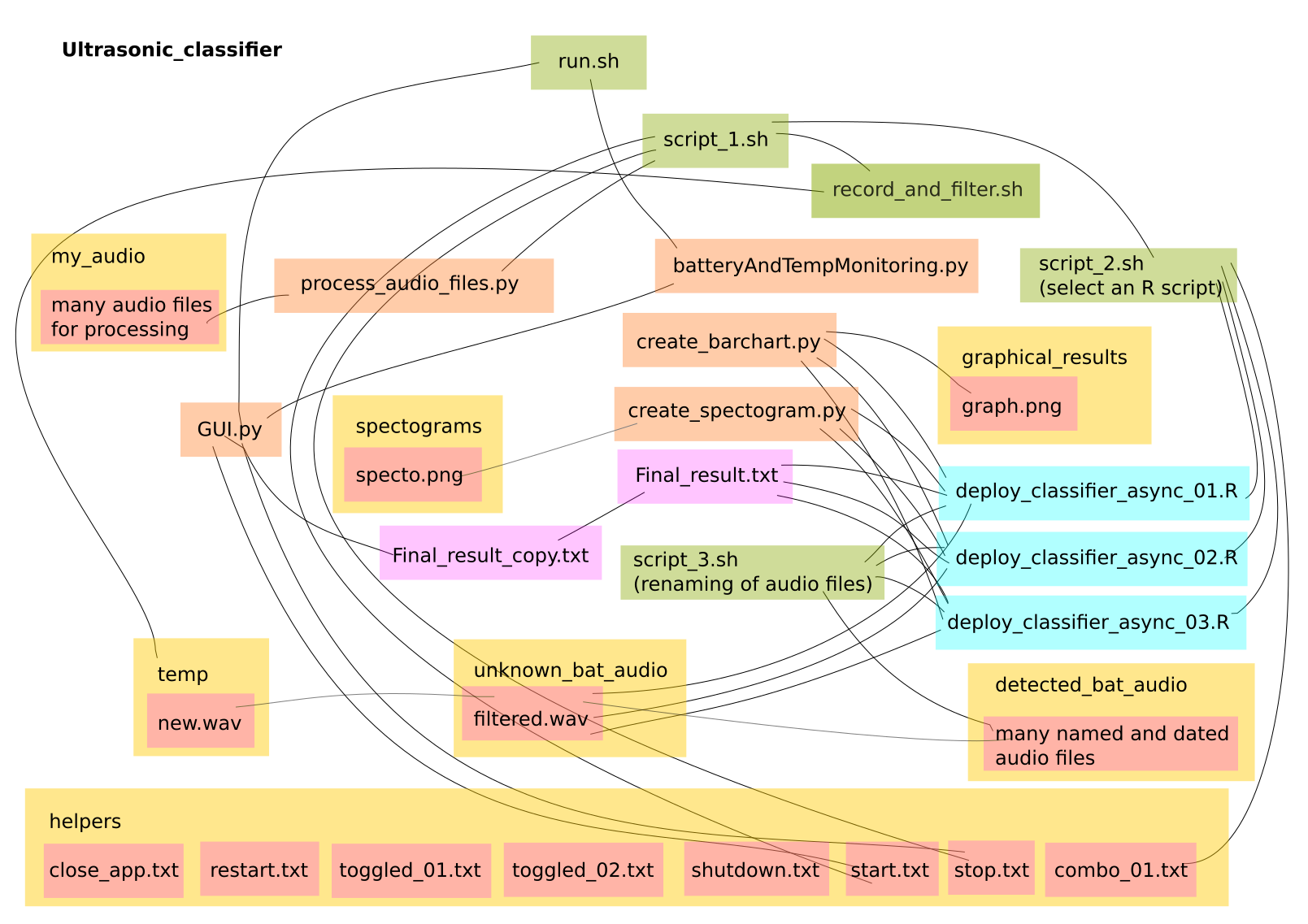
- Designing the overall architecture of the app - GTK only covers a very small part of the app - the touch screen display. The rest of it relies on various Bash and Python scripts to interact with the main deployment script which is written in R. Learning the R language was really not a problem as it's a very generic languages and and only seems to differ in it's idiosyncratic use of syntax, just like any other language really. The 'stack' architecture initially started to evolve organically with a lot of trial and error. As a Hacker, I just put it together in a way that seemed logical and did not involve too much work. I'm far too lazy to learn how to build a stack properly or even learn any language properly, but, after giving a presentation to my local university computer department, everybody seemed to agree that that was perfectly ok for product development. Below is a quick sketch of the stack interactions, which will be pure nonsense to most people but is invaluable to remind myself of how it all works:
![]()
- Creating a dynamic barchart - I really wanted to display the results of the bat detection system in the most easy and comprehensive way and the boring old barchart seemed like the way forwards. However, to make it a bit more exciting, I decided to have it update dynamically so that as soon as a bat...
 Capt. Flatus O'Flaherty ☠
Capt. Flatus O'Flaherty ☠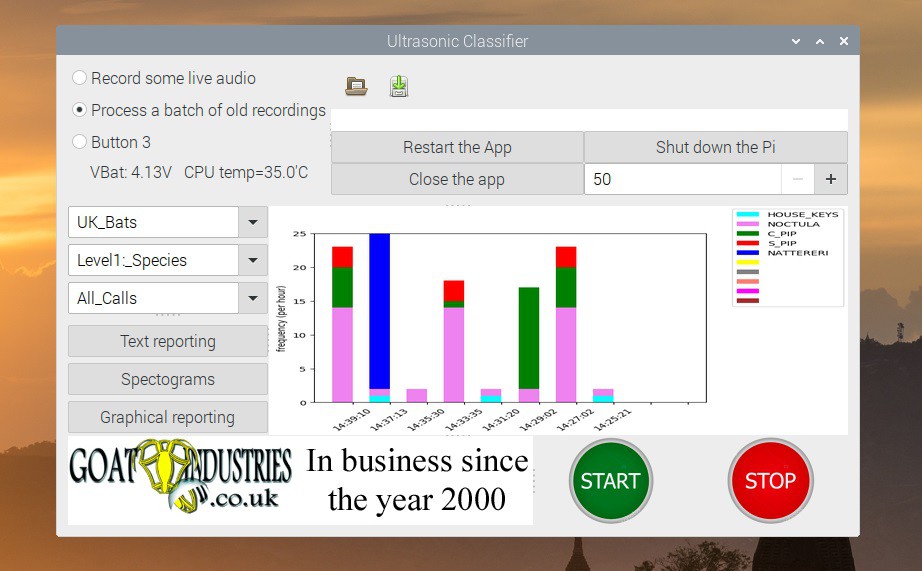

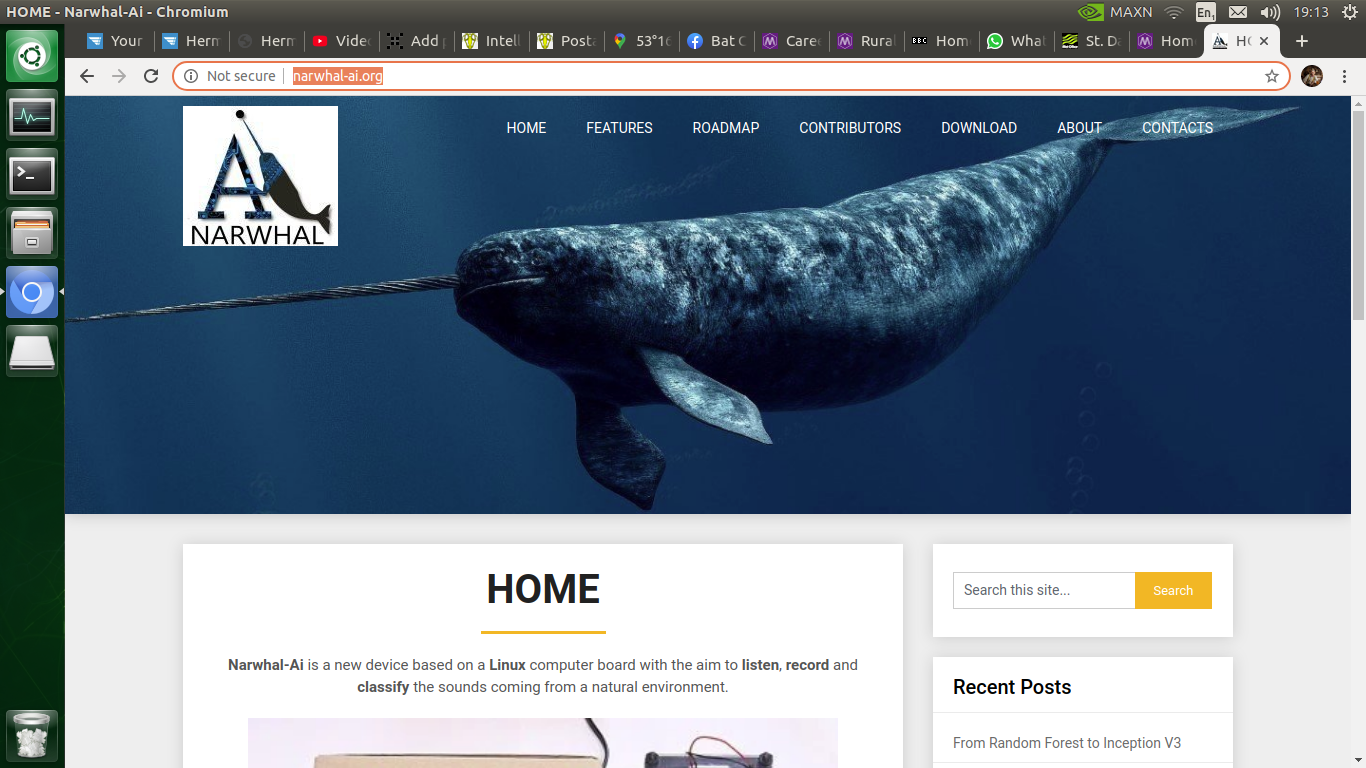



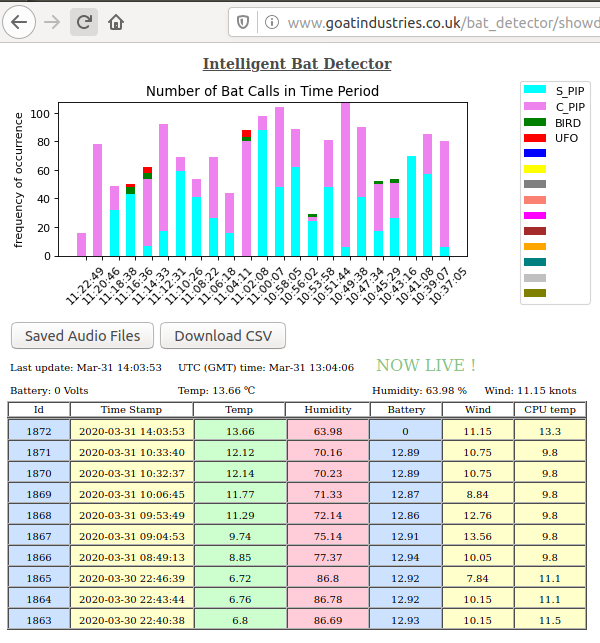 Most evenings, currently at about 20.00 UTC, this gadget will be deployed in the wild for rigorous testing and debugging. The barchart updates itself every 3 minutes or so and it's quite fun to see the animals appear during the evening / night.
Most evenings, currently at about 20.00 UTC, this gadget will be deployed in the wild for rigorous testing and debugging. The barchart updates itself every 3 minutes or so and it's quite fun to see the animals appear during the evening / night.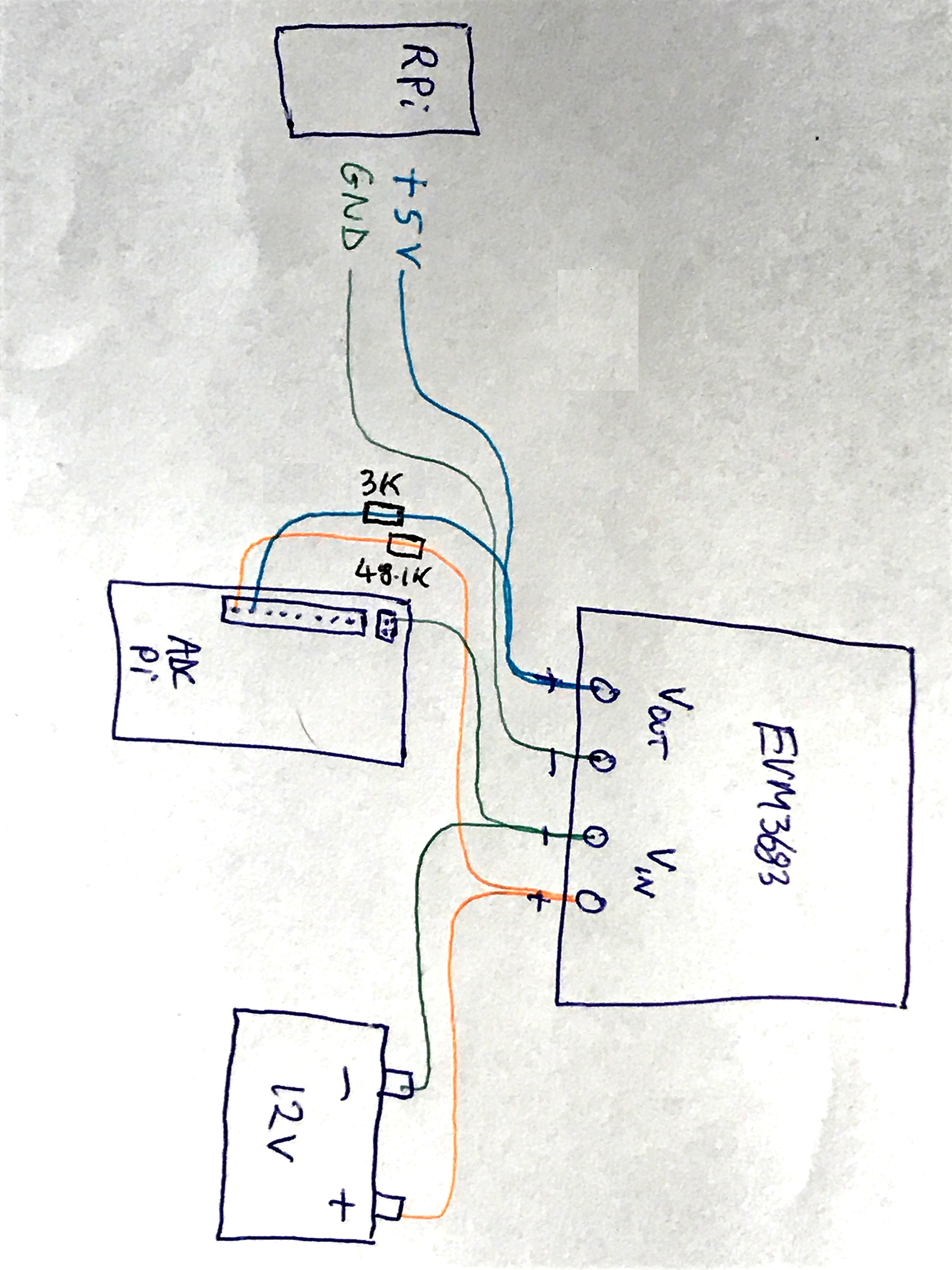






 John Grant
John Grant
 Rusty Jehangir
Rusty Jehangir
 Ashwin K Whitchurch
Ashwin K Whitchurch
 mick
mick
Can it be made cheaper? Eg with cheaper microphone and less computing force? It's not important to know right away what kind of bat it is right? Maybe data could be sent to the cloud to analyze?
There is a market for your product at the big windturbineproducers. In EU there are regulations on bats, and as we speak experiments are being done with bat detectors in windturbines to stall them when bats are flying out. You are always welcome to text us if you need more info.